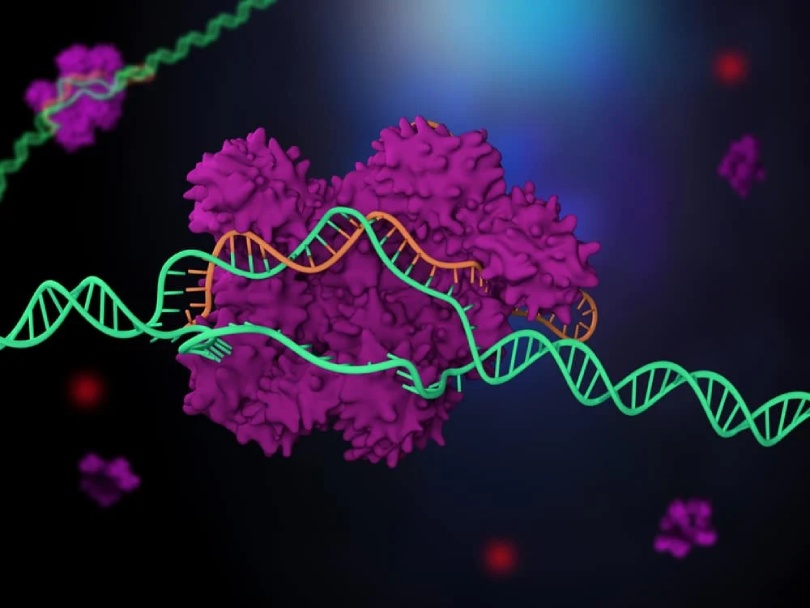
Specialists have distinguished another quality altering framework like CRISPR in complex creatures, showing interestingly that DNA-changing proteins exist across all realms of life.
Feng Zhang, a natural chemist at the Expansive Foundation of MIT and Harvard and the McGovern Establishment for Cerebrum Exploration at MIT, drove the group and already co-found the quality altering capability of the CRISPR-Cas9 framework, what capabilities as a sort of “sub-atomic scissors” that eliminate segments of DNA, in this way impairing qualities or permitting new ones to be traded in.
Until this discovery, these systems were only found in basic organisms like bacteria and archaea, which use them as a kind of primitive immune system to cut up invaders’ DNA. The new system, which is called Fanzor, was found in fungi, algae, amoebas, and a clam species. This significantly increased the number of groups that are known to use these genetic tools.
Ethan Bier, a geneticist at the University of California, San Diego, who uses gene editing in his work but was not involved in the study, stated, “People have been saying with such certainty for so long that eukaryotes [organisms whose complex cells contain nuclei] couldn’t have a similar system.” Be that as it may, it’s common shrewdness from the Zhang lab, discrediting them,” Casket told Live Science.
In the wake of distributing their most memorable paper on CRISPR in 2013, Zhang and associates started concentrating on how these frameworks advance. During this work, the gathering recognized a class of proteins in microbes called OMEGAs, remembered to be early predecessors of Cas9 proteins, the “scissors” of the CRISPR framework. They started to think that Fanzor proteins, a kind of OMEGA, could likewise be changing DNA.
The gathering evaluated web-based data sets for the proteins and were astounded to find a few in examples confined from growths, protists, arthropods, plants and goliath infections. According to Zhang, the hypothesis is that a process known as horizontal gene transfer occurred when the genes needed to make Fanzor proteins were transferred from bacteria to complex organisms. Eukaryotic organisms’ genomes contain transposable elements, or DNA fragments that are able to self-replicate and move around the genome. These elements contain the genes that encode for the Fanzor proteins.
In tests, the specialists found that Fanzor proteins share a few similitudes with CRISPR. In addition, fanzor proteins interact with guide RNA, a molecule that directs the proteins to the cut DNA. The strand of target DNA is enhanced by this molecule, which is called an omegaRNA. The two pieces zip together when they match, allowing Fanzor to cut the DNA.
The Fanzor system was tested in human cells by the team. At first, they found that it was only able to successfully add or remove bits of DNA about 12% of the time. However, the researchers were able to increase the efficiency to just over 18 percent after employing inventive engineering to improve and stabilize the system.
This failure isn’t is to be expected, as indicated by Casket, nor a sign that Fanzor isn’t comparable to CRISPR. Although “it certainly didn’t start out that way,” he stated, scientists have programmed CRISPR to make the desired substitutions almost always. However, Bier stated that Fanzor will have a difficult time competing with Cas9, which he described as “the most adaptable and forgiving protein for the types of things you want to do to it.”
Instead, it is likely that Fanzor will work in conjunction with CRISPR, which has been utilized in both research and experimental medical treatments for diseases like cancer and blindness.
Zhang stated that “the Fanzor systems are more compact and therefore have the potential to be more easily delivered to cells and tissues” in comparison to CRISPR. Additionally, they are less likely to degrade nearby RNA or DNA accidentally, which are known as “off-target” or “collateral effects.” As a result, gene therapy applications of Fanzor are appealing.
Zhang told Live Science he’s presently eager to go searching for comparable frameworks in new spots.
Zhang stated, “The power of studying biodiversity is really emphasized by this work.” There are probably many more natural RNA-guided systems that have the potential to edit genes in the future.